Biography
I am a computational chemist who builds predictive tools and studies reaction mechanisms using DFT, molecular dynamics, QM/MM, cheminformatics, and graph neural networks. I recently (February 2026) completed my Ph.D. with Prof. Peng Liu at the University of Pittsburgh. My work focuses on three main areas:
Transition-Metal and Enzyme Catalysis
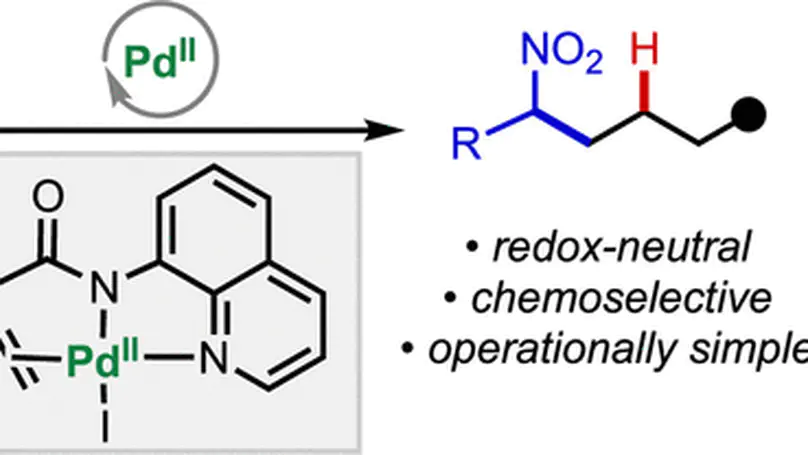
- Mechanistic Insights and Reaction Design Led computational studies on Ni/Pd/Cu/Rh-catalyzed reactions in collaboration with groups at Scripps (Keary Engle), Emory (Huw Davies), UC Davis (Annaliese Franz), UT Dallas (Vladimir Gevorgyan), and more, identifying catalyst and ligand features governing reactivity and selectivity.
- Enzyme Design DFT/MD work with the DeGrado (UCSF) and Yang (UCSB) labs on de novo designed metalloproteins. Predicted conformer-controlled selectivity in Si-H insertion and guided directed evolution to >99:1 er for Ge-H insertion (Science 2025, JACS 2025).
Cheminformatics and Machine Learning
- HArD (HeteroAryl Descriptors) Built a database of >31,500 monosubstituted heteroarenes with 65 descriptors (heteroaryl Hammett parameters, aromaticity metrics, HOMO-LUMO gaps, buried volumes). Published in Sci. Data 2025. You can try the tool here: hard.pengliugroup.com
- RAPID (Radical Polarity Predictor) Trained a graph neural network on >1 million DFT calculations to predict radical polarity for any organic radical from a drawn structure. Collaboration with the Nagib and Hutchison groups. Manuscript in preparation. You can try the tool here: radicalpolarity.pengliugroup.com
High-throughput MD simulations (on-going)
- Enzyme Selectivity Prediction Exploring whether physics-based descriptors from short classical MD simulations can enhance selectivity prediction for ~900 enzyme variants, building on the MODIFY framework for ML-guided directed evolution.
- Computational Chemistry (DFT, MD, QM/MM)
- Cheminformatics & Molecular Descriptors
- Machine Learning (GNNs, Property Prediction)
- Transition Metal & Enzyme Catalysis
Ph.D. in Chemistry, 2026
University of Pittsburgh
B.S. in Chemistry, 2021
Colorado State University
Publications
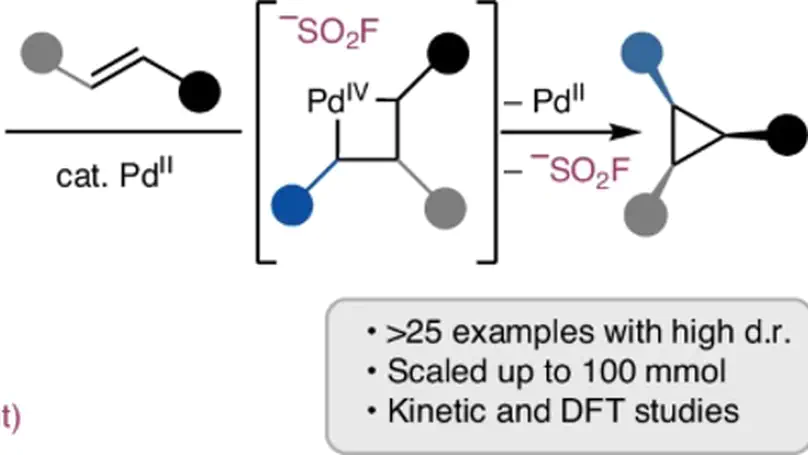
Alkyl sulfonyl fluorides act as ambiphilic coupling partners in Pd(II)-catalyzed stereoselective cyclopropanation of unactivated alkenes, with the SN2-type C–SO2F oxidative addition serving as the turnover-limiting and diastereoselectivity-determining step.
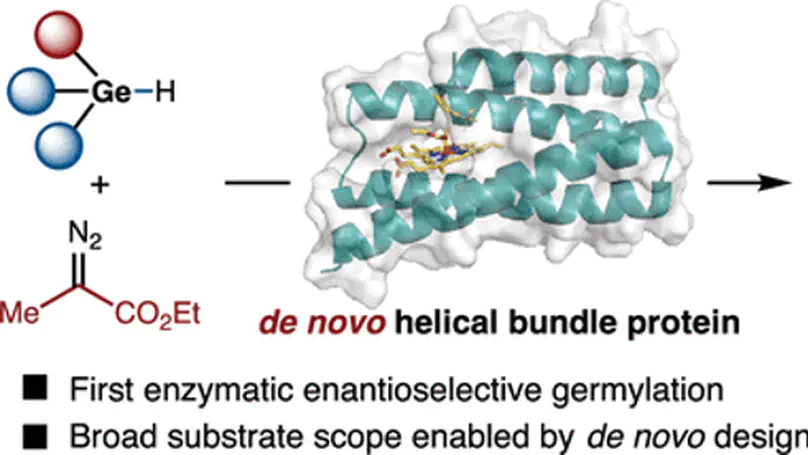
De novo designed helical bundle proteins were evolved to achieve enantioselective Ge–H insertion, the first enzymatic germylation reaction, with DFT and MD simulations revealing why Ge–H is mechanistically harder to control stereoselectively than Si–H insertion.
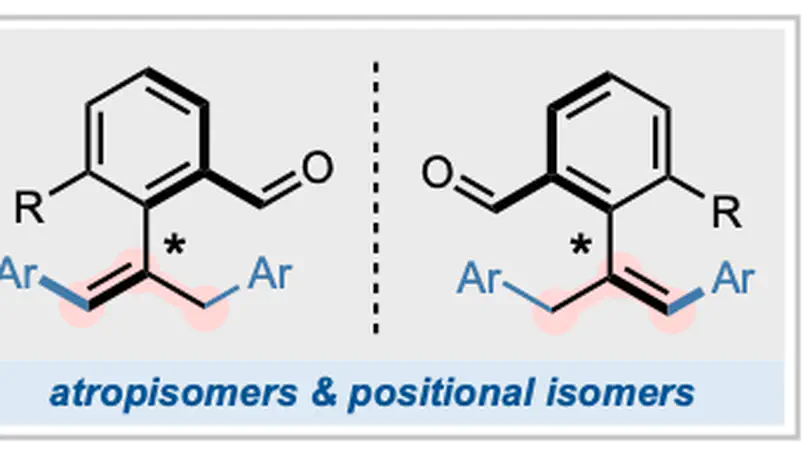
A Pd/amino acid cocatalytic system converts ortho-alkenyl benzaldehydes into styrene derivatives exhibiting the rare phenomenon of equivalent atrop- and positional isomerism via sequential enantioselective Mizoroki–Heck 1,3-diarylation.
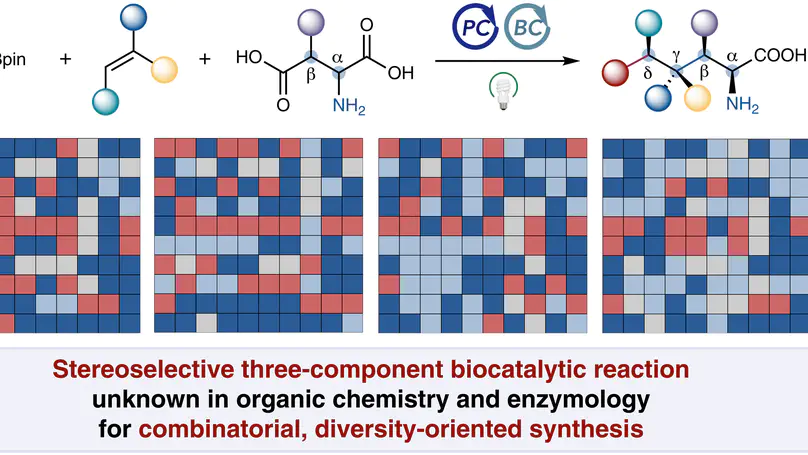
Directed evolution of pyridoxal decarboxylases enabled a stereoselective three-component radical C–C coupling new to both biochemistry and organic chemistry, providing six product classes with excellent stereocontrol for diversity-oriented medicinal chemistry library synthesis.
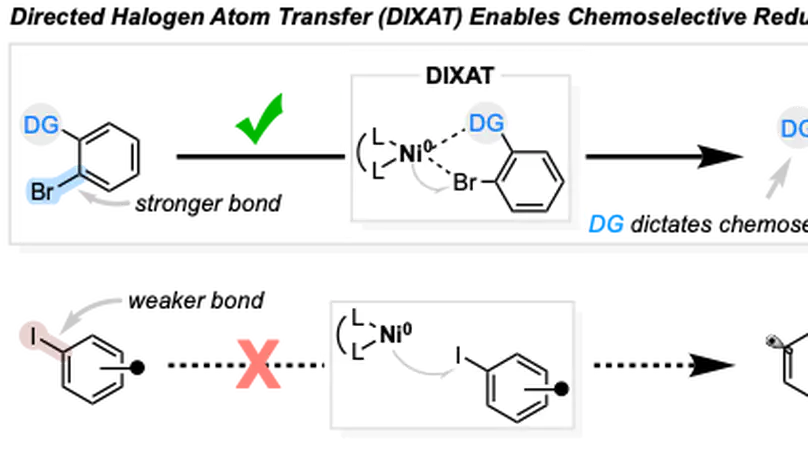
DIXAT (Directed Halogen Atom Transfer) is a new strategy that enables chemoselective activation of aryl bromides over aryl iodides through geometric directing effects, overturning the intrinsic reactivity trends of XAT and SET for Ni-catalyzed remote C(sp3)–H functionalization of aliphatic amines.
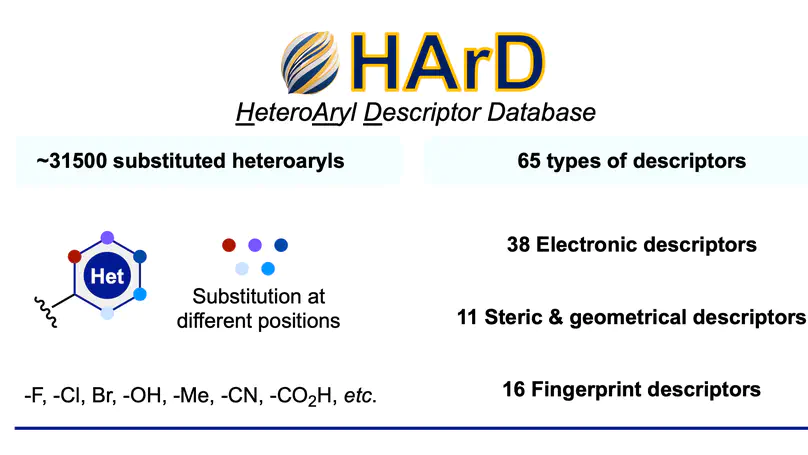

HArD is a freely accessible database of 65 DFT-computed steric and electronic descriptors for over 31,500 heteroaryl substituents, including a new Hammett-type substituent constant (σHet), designed to enable quantitative structure–activity and reactivity modeling of heteroaromatic compounds.
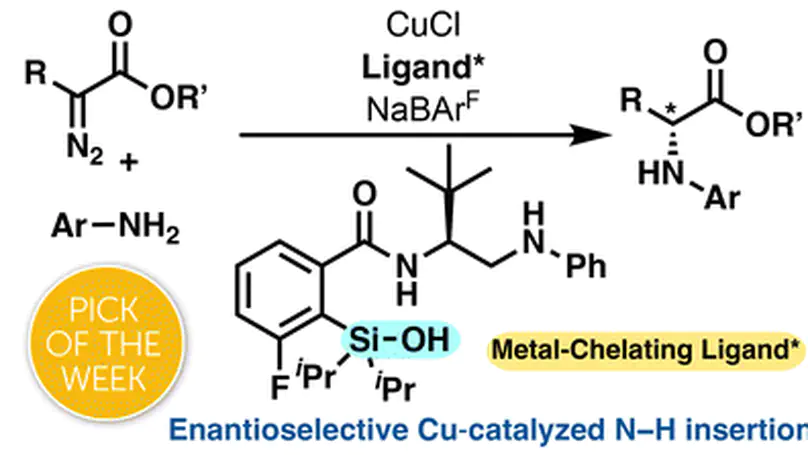
Novel chiral silanol ligands with a peptide-like aminoamide scaffold enable Cu-catalyzed enantioselective N–H insertion to form unnatural amino acids, with DFT and X-ray analysis revealing that H-bond-stabilized silanol–copper chelation and π–π stacking drive high selectivity.
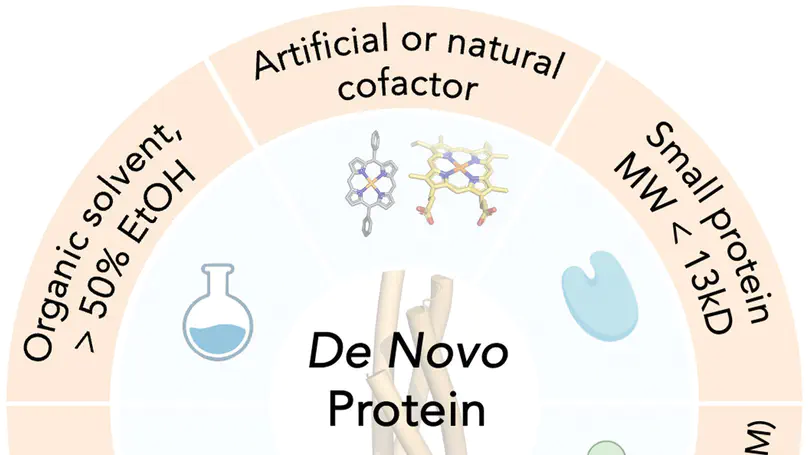
A de novo designed minimal four-helix bundle protein incorporating synthetic porphyrin or heme cofactors achieves high efficiency and enantioselectivity in abiological carbene transfer reactions, with directed evolution and computational analysis revealing the structural basis for stereocontrol.
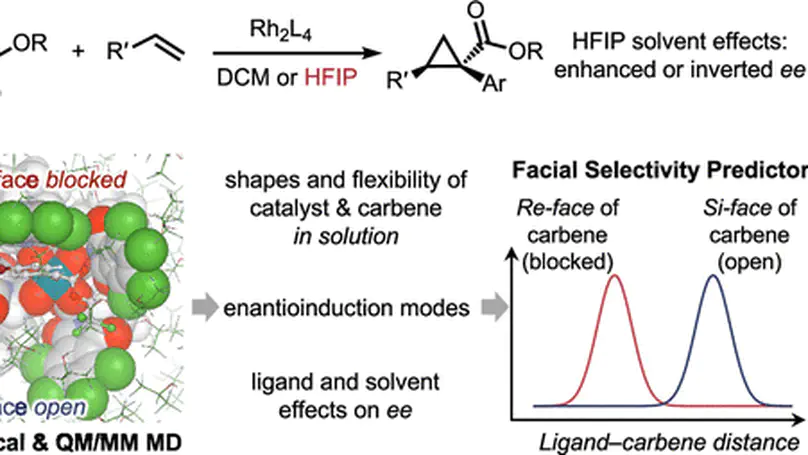
A combined experimental and computational study revealing how HFIP solvent induces flexibility in dirhodium catalysts to modulate enantioselectivity in cyclopropanation reactions.
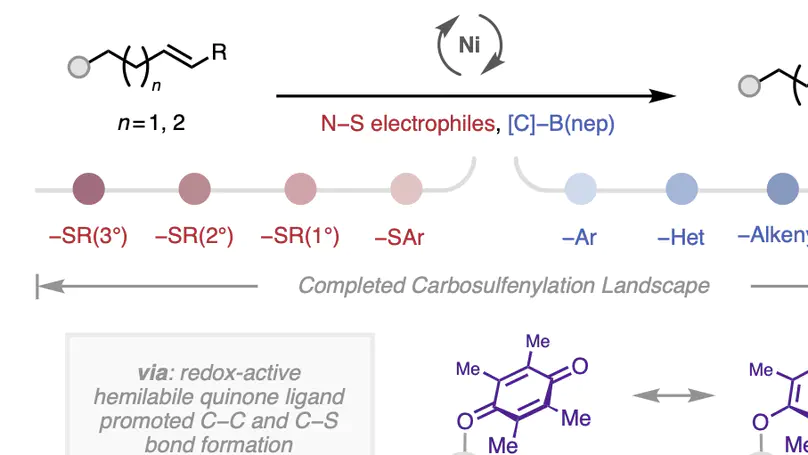
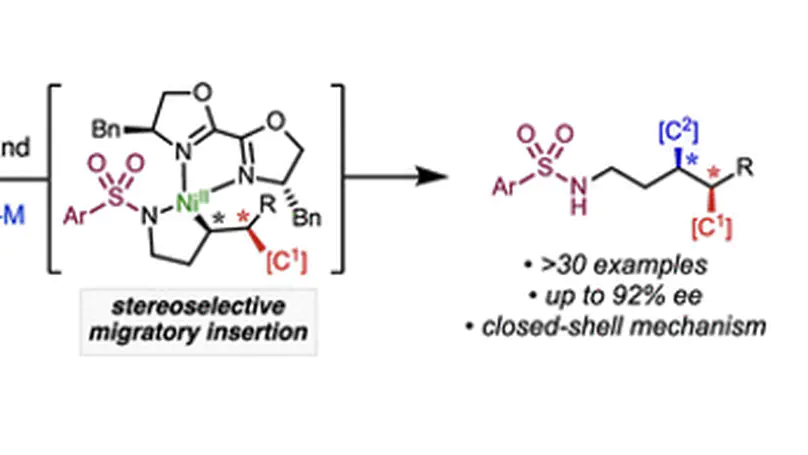
A three-component coupling approach towards structurally complex dialkylsulfides is described via the nickel-catalyzed 1,2-carbosulfenylation of unactivated alkenes with organoboron nucleophiles and alkylsulfenamide (N–S) electrophiles
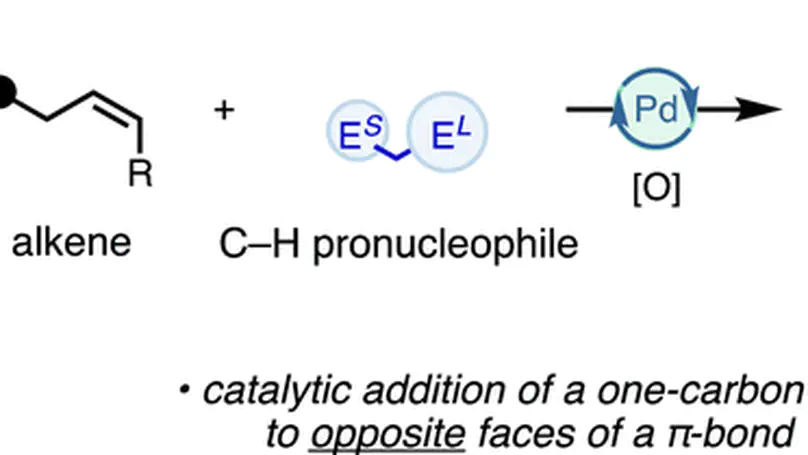
This study presents a mild and efficient method for synthesizing densely functionalized cyclopropanes via directed nucleopalladation of nonconjugated alkenes, yielding excellent anti-selectivity and accommodating diverse electron-withdrawing pronucleophiles, with mechanistic insights supported by DFT calculations showing key steps of α-iodination, anti-carbopalladation, and strain-release-promoted reductive elimination.
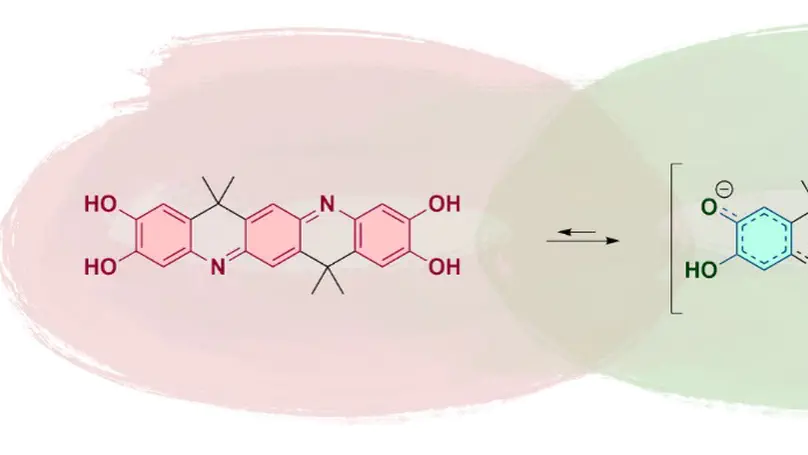
A novel resonance-assisted self-doping mechanism in ladder-type oligoaniline-derived organic conductors, enabling efficient proton transfer and enhanced stability without external dopants, as confirmed by mechanistic and computational studies.
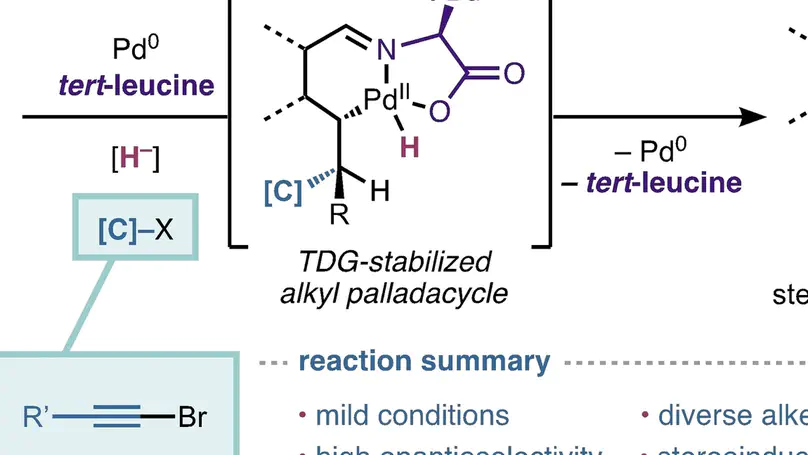
Herein, we report a transient directing group (TDG) strategy to facilitate site-selective palladium-catalyzed reductive Heck-type hydroalkenylation and hydroalkynylation of alkenylaldehyes using alkenyl and alkynyl bromides, respectively, allowing for construction of a stereocenter at the δ-position with respect to the aldehyde.
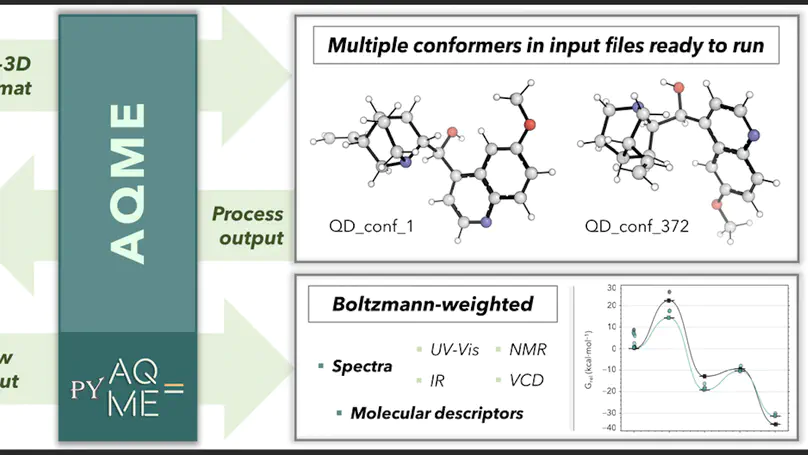
AQME, Automated Quantum Mechanical Environments, is a free and open-source Python package for the rapid deployment of automated workflows using cheminformatics and quantum chemistry.